Founded in 08/2015, the group is focused on biologically relevant molecules and macromolecular complexes, where intermoleculer interactions play a pivotal role in manifesting biological function. We are particularly interested in molecular level understanding of structure, function and mechanisms of membrane active compounds. Our aim is to study and design natural and nature-mimicking species which can be further developed in biomedical applications targeting organism-specific membranes.
Membrane targeting peptidic assemblies
The group focuses its research activities on molecular level understanding of structure, function and mechanisms of membrane active compounds, primarily natural and non-natural peptides with antimicrobial potency.
Natural antimicrobial peptides or host defence peptides constitute a part of the innate immune system of the host organisms and help in the fight against invading pathogens. They have a broad-spectrum activity against various bacteria, and fungi, in addition, they also show antiviral and anticancer properties. Their mechanism of action involves disruption of target cell membranes, moreover, they can elicit immunoregulatory actions as well. They represent a promising novel tool to combat multidrug-resistant bacteria, the emerging number of which represents a global threat. Beyond their applicability in human health care, antimicrobial peptides have potential in various fields, e.g. as biopesticide components to protect cultured plants in agriculture, nutrient additives in livestock, food preservatives in food industry, or even in wastewater treatment.
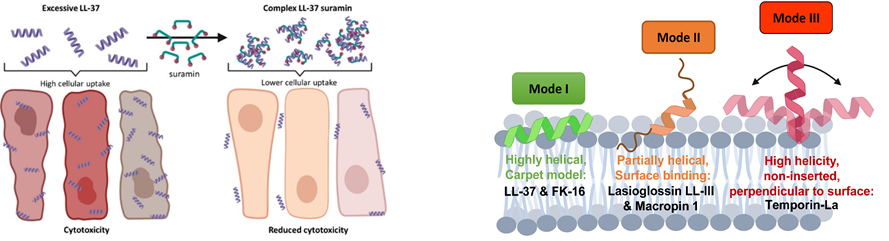
Despite the many advantages of natural antimicrobial peptides over common antibiotics, their proteolytic sensitivity can limit their medical use. In this regard, peptidomimetic agents called foldamers with structural and folding properties comparable to their natural templates can represent a good alternative. Non-natural peptides display excellent self-organising properties ranging from toxic amyloid oligomers to ion channel formation, with tunable applications ranging from tissue engineering through antiviral and antibacterial therapeutics to supramolecular polymer tubes.

Selected related publications:
- Juhász et al “Interplay Between Membrane Active Host Defense Peptides And Heme Modulates Their Assemblies And In Vitro Activity” (2021) SciRep
- Quemé-Peña et al “Membrane Association Modes Of Natural Anticancer Peptides: Mechanistic Details On Helicity, Orientation, And Surface Coverage” (2021) IJMS
- Zsila et al “Quorum Sensing Pseudomonas Quinolone Signal Forms Chiral Supramolecular Assemblies With The Host Defense Peptide Ll-37” (2021) Front Mol Biosci
- Ricci et al “Anionic Food Color Tartrazine Enhances Antibacterial Efficacy Of Histatin-Derived Peptide Dhvar4 By Fine-Tuning Its Membrane Activity” (2020) QRB
- Szigyártó et al “Membrane Active Janus-Oligomers Of β ³-Peptides” (2020) Chem Sci
- Udyavara Nagaraj et al “Stimuli-Responsive Membrane Anchor Peptide Nanofoils for Tunable Membrane Association and Lipid Bilayer Fusion” (2022) ACS Appl Mater Interfaces
Molecular engineering of extracellular vesicles
We also focus on standardized isolation and characterization protocol development for extracellular vesicles and on the molecular engineering of these species.
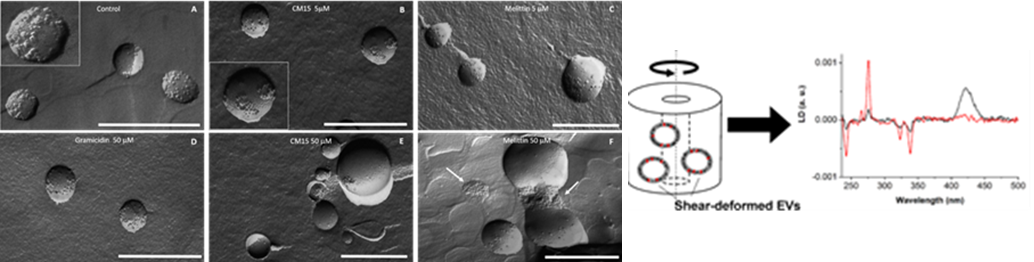
Extracellular vesicles (EVs) are biological nanoparticles, surrounded by a phospholipid bilayer and released by various cells into the extracellular space. They are classified based on their cellular origin, biogenesis and physicochemical properties. They can carry different type of biomolecules (such as DNA, RNA, proteins, phospholipids) which are characteristic to the particular cells producing them. Besides membrane inserted proteins, in vivo systems can also contain biomolecules adsorbed on their surface, known as protein corona. Evs are important mediators of intracellular communication and are involved in many physiological and pathological processes, so they may provide a wealth of information for early diagnosis and treatment of various diseases. Beyond these roles, they can be used as potential delivery vehicles for different drugs, food nutrient, inorganic nanoparticles etc. Therefore, the composition, modulation and characterization of protein corona is essential, as can influence their bioavailability, bio-distribution and their applicability.

Selected publications:
- Szigyártó et al., Flow-Alignment of Extracellular Vesicles: Structure and Orientation of Membrane Associated Biomacromolecules Studied with Polarized Light, 2018, ChemBioChem
- Singh et. al., Membrane Active Peptides Remove Surface Adsorbed Protein Corona from Extracellular Vesicles of Red Blood Cells, 2020, Frontiers in Chemistry
Computational membrane biology
Besides experimental investigations, we employ all-atom Molecular Dynamics (MD), Coarse-Grained MD as well as quantum chemical simulations with numerous antimicrobial peptides (AMP) in order to investigate their behaviour and binding conformations on complex membranes, extracellular vesicles (EVs), and single composition lipid bilayers. We are particularly interested in studiyng peptide interactions with small molecules (eg. with drug molecules, food colors, bacterial and human signalling compounds, etc.). Here it is crucial to understand how how these interactions can cause structural changes in AMPs as well as affect their membrane binding properties. Our studies are often supplemented with molecular Docking tools that helps identifying initial docking sites of the molecules on larger species and assists model building when experimental structural details are lacking. Furthermore simulations enable calculation of electrostatic potentials, entropy, binding free energy and other quantifiable parameters, which greatly helps for objective comparison of different systems. Lastly quantum chemical calculations are also frequenatly used in order to parametrize molecules for MD simulations and also to assess their photophysical and conformational properties.
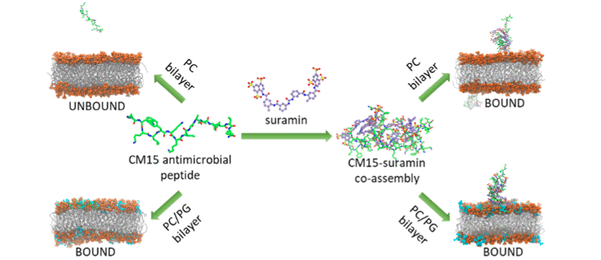
References:
- Kohut, Gergely, Tünde Juhász, Mayra Quemé-Peña, Szilvia Bosze, and Tamás Beke-Somfai. 2021. “Controlling Peptide Function By Directed Assembly Formation: Mechanistic Insights Using Multiscale Modeling On An Antimicrobial Peptide–Drug–Membrane System”. Acs Omega 6 (24). American Chemical Society (ACS): 15756-15769.
- Quemé-Peña, Mayra, Tünde Juhász, Gergely Kohut, Maria Ricci, Priyanka Singh, Imola Cs. Szigyártó, Zita I. Papp, Lívia Fülöp, and Tamás Beke-Somfai.(01/2021AD) 2021. “Membrane Association Modes Of Natural Anticancer Peptides: Mechanistic Details On Helicity, Orientation, And Surface Coverage”. International Journal Of Molecular Sciences 22 (16).
- Szigyártó, Imola Cs., Judith Mihály, András Wacha, Dóra Bogdán, Tünde Juhász, Gergely Kohut, Gitta Schlosser, et al. 2020. “Membrane Active Janus-Oligomers Of Β ³ -Peptides”. Chemical Science 11: 6868–6881
- Pothoff, Jan, Krzysztof Kamil Bojarski, Gergely Kohut, Agnieszka G Lipska, Adam Liwo, Efrat Kessler, Sylvie Ricard-Blum, and Sergey A. Samsonov. 2019.
“Analysis Of Procollagen C-Proteinase Enhancer-1/Glycosaminoglycan Binding Sites And Of The Potential Role Of Calcium Ions In The Interaction”. International Journal Of Molecular Sciences 20 (20). - Zsila, Ferenc, Gergely Kohut, and Tamás Beke-Somfai. (05/2019AD) 2019.
“Disorder-To-Helix Conformational Conversion Of The Human Immunomodulatory Peptide Ll-37 Induced By Antiinflammatory Drugs, Food Dyes And Some Metabolites”. International Journal Of Biological Macromolecules 129: 50-60. - Kohut, Gergely, Adam Sieradzan, Ferenc Zsila, Tünde Juhász, Szilvia Bosze, Adam Liwo, Sergey A. Samsonov, and Tamás Beke-Somfai. 2019.
“The Molecular Mechanism Of Structural Changes In The Antimicrobial Peptide Cm15 Upon Complex Formation With Drug Molecule Suramin: A Computational Analysis”. Physical Chemistry Chemical Physics 21: 10644–10659. - Kohut, Gergely, Adam Liwo, Szilvia Bosze, Tamás Beke-Somfai, and Sergey A. Samsonov. 2018.
“Protein-Ligand Interaction Energy-Based Entropy Calculations: Fundamental Challenges For Flexible Systems”. The Journal Of Physical Chemistry B.
- Circular and Linear dichroism
- UV/Vis absorption spectroscopy
- Computational biophysics
- Sum-frequency generation spectroscopy
- Supported lipid bilayers
- Liposomal systems
- Bicelles













